Effects of knockdown of autophagy pathway genes onlongevity are highly condition dependent

专属客服号

微信订阅号
大数据治理
全面提升数据价值
赋能业务提质增效
Abstract
Autophagy is proposed to protect against aging by clearing damaged cellular constituents. In line with this several life-extending interventions in model organisms show some degree of autophagy dependence. In
C. elegans
, inhibiting autophagy can shorten, lengthen or have no effect on lifespan. Differences between published findings likely reflect variability in experimental conditions. Here we investigate the condition dependence of effects on lifespan of RNA-mediated interference (RNAi) knockdown of autophagy pathway components. Effects on several interventions causing a strong Age (increased lifespan) phenotype were examined, including mutation of
daf-2
(insulin/IGF-1 receptor). Factors varied included
daf-2
mutant allele class,
atg
gene, temperature and presence of 5-fluoro-2’-deoxyuridine (FUDR). Effects on lifespan of
atg
RNAi proved to be highly condition dependent. Notably, for most
atg
genes tested lifespan was not usually reduced more in the long-lived mutant than in the wild-type control. Greater suppression was seen at 20°C for certain
atg
genes with
daf-2(e1368)
but not
daf-2(e1370)
. At 25°C, little reduction in lifespan was seen. However,
atg-18
knockdown behaved differently, suppressing
daf-2
Age under all conditions, suggesting possible pleiotropic action. FUDR at a high concentration caused knockdown of several
atg
genes to increase lifespan. Thus, depending on experimental conditions,
atg
knockdown can increase, decrease or have no effect on
daf-2
Age. The lack of suppression of Age by
atg
RNAi under most conditions questions the importance of autophagy in
daf-2
Age. Moreover, condition dependence of effects creates a risk of possible condition selection bias.
Introduction
A long-standing theory is that senescence (aging) is largely a consequence of the accumulation of random molecular damage caused by, among other things, reactive oxygen species (
Beckman and Ames, 1998
;
Harman, 1956
;
Murphy, 2023
;
Shore and Ruvkun, 2013
;
Szilard, 1959
). This view predicts that mechanisms of somatic maintenance, particularly those that prevent accumulation of damaged cellular constituents, will decelerate the aging process. One somatic maintenance function viewed as a potential longevity-assurance process is autophagy (specifically macroautophagy), which effects lysosome-dependent degradation of cellular constituents, including damaged matter (
Aman et al., 2021
).
The possible anti-aging role of autophagy has been extensively tested in the short-lived nematode
Caenorhabditis elegans
, and supporting evidence found with respect to several life-extending interventions, including reduced insulin/IGF-1 signaling (IIS), reduced germline signaling, and dietary restriction (
Hansen et al., 2018
). These studies were principally performed in the 2000s; however, during the same period, falsification tests of the molecular damage theory, particularly that relating to oxidative damage, led to some uncertainty about its validity (
Gems and Doonan, 2009
;
Perez et al., 2009
;
Shields et al., 2021
). Meanwhile, an alternative theoretical framework emerged, based on the evolutionary theory of aging (
Arnold and Rose, 2023
;
Williams, 1957
), arguing that senescence is largely the consequence of genetically-determined, programmatic mechanisms (
Blagosklonny, 2006
;
de Magalhães and Church, 2005
;
Gems, 2022
;
Maklakov and Chapman, 2019
), and very much so in
C. elegan
s (
Gems and de la Guardia, 2013
;
Gems et al., 2021
;
Pires da Silva et al., 2024
).
One form of programmatic aging involves costly programs: genetically-determined processes that degrade somatic tissues as a by-product of wider, fitness-promoting processes (
Gems and Kern, 2024
;
Gems et al., 2021
). In some cases this involves biomass repurposing, where biomass of one tissue is broken down by autophagic processes to release molecular constituents to support functions in another. This occurs to a high degree in semelparous organisms in the process of reproductive death (suicidal reproductive effort), as in many monocarpic plants, and semelparous fish such as Pacific salmon (
Gems et al., 2021
).
Several lines of evidence support the hypothesis that reproductive death occurs in
C. elegans
hermaphrodites (
Gems et al., 2021
;
Kern et al., 2023
). This includes a putative costly program in which intestinal biomass is broken down and repurposed to support production of yolk, that is then vented to support larval growth, leading to intestinal atrophy (a senescent pathology) (
Ezcurra et al., 2018
;
Kern et al., 2021
;
Sornda et al., 2019
). Notably, such biomass repurposing is supported by autophagy, as evidenced by deceleration of intestinal atrophy and yolk pool formation when autophagy is inhibited (
Benedetto and Gems, 2019
;
Ezcurra et al., 2018
). Thus, in this particular context autophagy appears to be promoting rather than preventing senescence.
These developments warrant a careful re-examination of the evidence that autophagy is protective against aging in
C. elegans
. In this study we re-examine the question of whether the longevity of
daf-2
insulin/IGF-1 receptor mutants is autophagy dependent. Here the principal form of past evidence involves epistasis tests, where effects on lifespan of reduction of function of genes encoding proteins effecting autophagy is compared in the wild type (N2) and
daf-2
mutants. Findings from 8 previous studies involving 46 epistasis experiments are summarised in
Table S1
. Although it is widely believed that autophagy is essential for
daf-2
mutant longevity (
Meléndez et al., 2003
), scrutiny of the results of past tests raises doubts about the strength of this claim.
Careful consideration of these prior studies identifies six distinct limitations, as follows. (1) A life-shortening effect of inhibition of autophagy does not necessarily indicate its role in the normal aging process, or in
daf-2
mutant longevity. (2) If autophagy is inhibited during development as well as adulthood, a life-shortening effect could result from disruption of normal development. (3) The claim that
daf-2
longevity is autophagy dependent requires evidence that the life-shortening effect in
daf-2
is significantly greater than in wild type, e.g. using Cox proportional hazard analysis, and this is rarely performed. (4) If effects of reducing autophagy are condition dependent, this introduces a potential bias: a risk that investigators might unwittingly select conditions where the data generated supports a role of autophagy in longevity - what may be referred to as
condition selection bias
. Potential determinative conditions that have varied across studies include choice of autophagy-determining gene to inhibit, of
daf-2
mutant allele, and ambient temperature. (5) 5-fluoro-2’-deoxyuridine (FUDR) is sometimes used to facilitate
C. elegans
lifespan assays by preventing progeny hatching; this could potentially affect test outcomes, as shown in other contexts (
Aitlhadj and Sturzenbaum, 2010
;
Anderson et al., 2016
;
Burnaevskiy et al., 2018
;
Van Raamsdonk and Hekimi, 2011
;
Zhao et al., 2019
). (6) A final, straightforward issue is whether a given finding proves to be reproducible in subsequent reports under, seemingly, the same conditions.
Of 46 prior tests (
Table S1
) only 4 present clear evidence that reducing autophagy shortens lifespan more in
daf-2
than in the wild type. Regarding one of these four instances, the effect of
bec-1
RNAi on
daf-2(e1370)
at 15°C (
Meléndez et al., 2003
), a subsequent study did not replicate this finding (
Hars et al., 2007
). In two other cases the weaker
daf-2(mu150)
allele was used (
Hansen et al., 2008
;
Patel et al., 2008
). In two further studies where large reductions in
daf-2
lifespan were observed (
Chang et al., 2017
;
Minnerly et al., 2017
) knockdown of
atg-18
was used. In a 2009 study where effects of knockdown of 14 different autophagy genes was tested, adult-limited
atg-18
RNAi was one of only 2/14 that significantly reduced lifespan in the wild type (
Hashimoto et al., 2009
), suggesting possible
atg-18
idiosyncrasy. The 2009 study includes more than half of all of the prior tests (28/46); strikingly, in only 1/14 genes (
atg-4.1
) did adult-limited RNAi significantly reduce
daf-2(e1370)
lifespan, while for 3/14 genes (
atg-9
,
bec-1
and
unc-51
) it
increased daf-2
lifespan (
Hashimoto et al., 2009
). One possible reason for the lack of observed life-shortening effects given autophagy knockdown is its use of FUDR at a high concentration.
In this study we assess the condition-dependence of the effects of RNAi knockdown of autophagy genes on
C. elegans
longevity. To this end we have tested the effects of RNAi of six genes in the autophagy pathway on longevity in two different
daf-2
mutants, and a
glp-1
germlineless mutant, at two temperatures. We have also assessed effects of FUDR and prevention of bacterial infection. Our results provide a robust foundation of evidence relating to possible autophagy dependence of
daf-2
and
glp-1
longevity. They suggest a weak and highly-condition dependent contribution of autophagy to
daf-2
and
glp-1
Age. This implies that prevention of damage accumulation by autophagy is, at most, a minor determinant of
daf-2
longevity.
Results
Effects of
atg
RNAi on
daf-2
Age vary with
daf-2
allele
We first tested whether effects on lifespan of inhibiting autophagy depend upon
daf-2
allele severity. Two
daf-2
mutants were examined.
daf-2(e1368)
is a class 1 (less pleiotropic) mutant where adults show normal behavior at both 20°C and 25°C, while
daf-2(e1370)
is a class 2 (more pleiotropic) mutant where at 25°C adults become paralyzed and cease feeding (
Gems et al., 1998
). Knockdown of autophagy was performed by RNAi from the L4 (late larval) stage, to exclude possible confounding, life-shortening disruption of normal development. RNAi was performed for six genes at several stages of the autophagy process: initiation (
atg-13
), membrane nucleation (
atg-9
,
bec-1
), phagophore formation (
atg-2
,
atg-18
) and elongation (
atg-4.1
) (
Figure 1A
). 5/6 genes were those examined in our previous study (
Ezcurra et al., 2018
), with
bec-1
added because it was the subject of several earlier studies (
Table S1
).
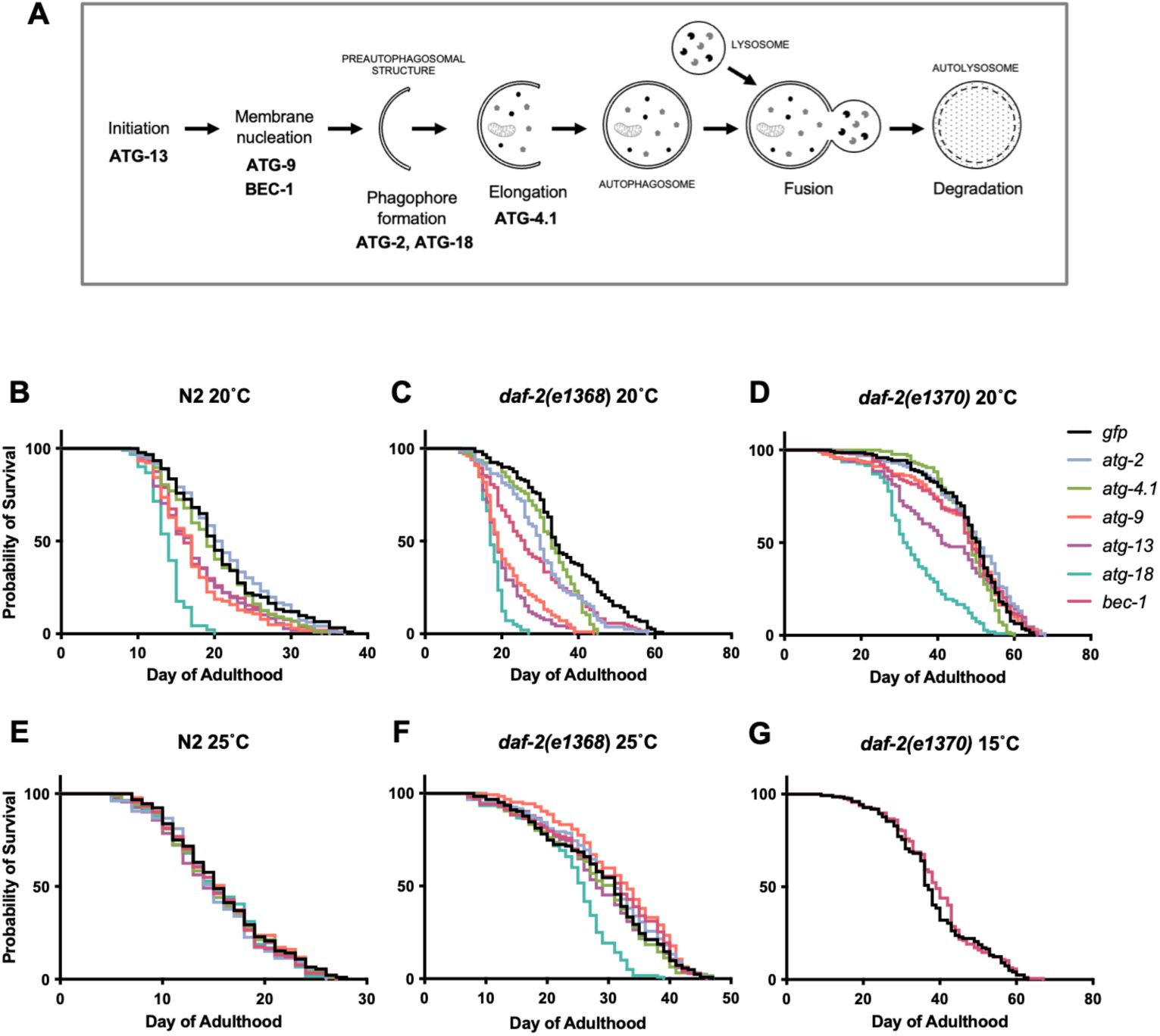
Effects of
atg
RNAi on longevity are highly variable and condition dependent.
(A) The autophagy pathway, and genes tested in this study. (B-D) Effects at 20°C. (B) Effects on N2 (wild type). (C) Effects on
daf-2(e1368)
. (D) Effects on
daf-2(e1370)
. (F, G) Effects at 25°C. (F) Effects on N2. (G) Effects on
daf-2(e1368)
. (H) Effects of
bec-1
RNAi on
daf-2(e1370)
at 15°C. (B-G summed data,
N
= 2; for individual trials and statistical comparisons, see
Table S2
,
S5
.
A methodological note: for tests of effects of a given intervention on
C. elegans
lifespan an often-applied standard is to include 3 biological replicates. This is true of several recent studies where the effect of knockdown of a single
atg
gene on
daf-2
longevity was studied (
Minnerly et al., 2017
;
Wilhelm et al., 2017
;
Yang et al., 2024
). However, given that the present condition dependence study effectively performs this test in 18 different ways, involving RNAi of 6
atg
genes, 2
daf-2
mutants and 2 temperatures,
N
= 2 biological replicates were judged to be sufficient to draw robust conclusions; similarly, an earlier study of RNAi 14
atg
genes under two conditions used 2-3 biological replicates (
Hashimoto et al., 2009
); for an overview of
N
sizes in previous studies, see
Table S1
.
Trials were performed at 20°C, and used
gfp
RNAi as a negative control. In wild-type
C. elegans
(N2), statistically significant reductions were sometimes seen:
atg-9, atg-13
and, particularly,
atg-18
RNAi shortened lifespan in both trials (summed data,
N
= 2, means: −13.7%,
p
< 0.0001, −12.4%,
p
= 0.0002, −31%,
p
< 0.0001, respectively; log rank test) (
Figure 1B
,
Table S2
; for all raw mortality data, see Supplemental Dataset 1). For survival curves comparing RNAi effects of individual
atg
genes on the three genotypes, see
Figure 2
.

Degree of suppression of
daf-2
longevity by
atg
gene RNAi differs greatly between
daf-2
alleles and
atg
RNAi treatments (20°C), cf.
Figure 1B-D
(same data).
Summed data,
N
= 2; for individual trials and statistical comparisons, see
Table S2
. All trials were performed in parallel, i.e. all lifespans are directly comparable.
RNAi of all six
atg
genes consistently reduced lifespan in the
daf-2(e1368)
mutant, with effects ranging from −10.5% (
p
= 0.0009) for
atg-4.1
to −50.4% (
p
< 0.0001) for
atg-18
(
Figure 1C
,
Table S2
). To assess whether
atg
RNAi reduced lifespan more in
daf-2
mutants than in the wild type, Cox proportional hazard (CPH) analysis was used. This showed a significantly greater effect in
daf-2(e1368)
for 4/6 genes (exceptions:
atg-4.1
,
bec-1
) (
Table S2
). These findings are in line with the earlier observation that
bec-1
and
vps-34
RNAi shortened the lifespan of the
daf-2(mu150)
class 1 mutant but not of N2 at 20°C (
Hansen et al., 2008
).
To test whether
atg
RNAi is able to fully suppress
daf-2(e1368)
Age, the lifespans of N2 and
daf-2
subjected to
atg
RNAi were compared. Under all six
atg
RNAi conditions,
daf-2(e1368)
still significantly increased mean lifespan, from +15.7% (
p
= 0.0031) for
atg-13
to +60.9% (
p
< 0.0001) for
atg-4.1
(summed data, 20°C,
Table S2
). This could imply either that
daf-2(e1368)
Age is not fully autophagy dependent, or that it is but autophagy is not fully suppressed by the RNAi treatment.
In
daf-2(e1370)
,
atg
RNAi had far more modest effects on lifespan than in
daf-2(e1368)
. Summed data showed significant reductions in lifespan resulting from RNAi of
atg-9
,
atg-13
and
atg-18
only, with the latter again causing the largest reduction: −8.3% (
p
= 0.012), −17.8% (
p
= 0.0034), and −29.2% (
p
< 0.0001), respectively (
Figure 1D
,
Table S2
). However, effects were in no instance significantly greater than in N2 (CPH analysis,
Table S2
), thus failing to provide evidence that
e1370
longevity is mediated by autophagy. Taken together, these results could imply that at 20°C autophagy contributes to longevity in weaker but not stronger
daf-2
mutants, possibly in class 1 but not class 2 mutants.
RNAi effects varied between
atg
genes, with
atg-2
and
atg-4.1
RNAi effects weaker, and
atg-18
RNAi effects generally stronger than the rest. One possible cause of this variability is differences between the plasmid vectors used for RNAi in terms of how efficiently they destroy their target mRNA. To investigate this possibility, RNAi of each
atg
gene was performed in N2 animals, and transcript levels measured by RT–qPCR and analyzed using the ΔΔCt method, normalized to a
gfp
RNAi control, with fold changes calculated as 2^−ΔΔCt (
n
= 4 independent trials). Effects of RNAi on mRNA level varied greatly, from no detected reduction in
atg-2
to an 86% reduction in
atg-18
(
Fig. S1A
;
Table S3
;
Supplementary dataset 2
). Pairwise comparisons of ΔΔCt values showed that,
atg-2
aside, there were no significant differences between RNAi treatment effects on mRNA levels, apart from a greater effect of
atg-18
RNAi when compared to either
atg-4.1
and
atg-9
RNAi (
Fig. S1B
).
Comparing effects of RNAi on mRNA levels and lifespan, the only clear correspondence between the former and the latter involved
atg-2
and
atg-4.1
RNAi, which in N2 had no significant effect on either (
Figure 1B
,
Fig. S1A
;
Table S2
). However for the remaining 4 genes, there was no clear correspondence, though
atg-18
mRNA levels appear to be the lowest, in line with its greater effect on lifespan (
Figure 1B-D
,
Fig. S1A
;
Table S2
,
Table S3
). These results imply that RNAi efficacy and perhaps also gene-specific issues contribute to variation in
atg
RNAi effects on lifespan (discussed further below). While mRNA levels after RNAi under the various other conditions tested were not assayed, reduced IIS (including
daf-2(e1370)
) intensifies the RNAi response (
Wang and Ruvkun, 2004
), thus lack of effect on lifespan in
daf-2(e1370)
is very unlikely to reflect suppression of mRNA knockdown.
Effect of
atg
RNAi on
daf-2
Age is temperature dependent
Results of a previous study performed at 25°C appear to show no greater reduction in lifespan in
daf-2(e1370)
compared to
daf-2(+)
after autophagy knockdown, even when using the
atg-18(gk378)
deletion mutation (
Toth et al., 2008
) (
Table S1
), suggesting possible temperature dependence of
atg
RNAi effects. To explore this further, in parallel to tests at 20°C, we also compared effects of
atg
RNAi on lifespan in N2 and
daf-2(e1368)
at 25°C. Effects on
daf-2(e1370)
were not tested, partly because this mutant ceases feeding at 25°C (
Gems et al., 1998
) which would be expected to interfere with RNAi by feeding.
At 25°C,
atg
RNAi did not shorten N2 lifespan for any of the six genes tested, not even
atg-18
(summed data;
Figure 1E
,
Table S2
), i.e. culture at 25°C suppressed the life-shortening effect of
atg
RNAi in N2. Also notable is that the increases in lifespan with
atg-2
and
atg-13
RNAi at 25°C, described in our earlier study (
Ezcurra et al., 2018
), were not reproduced (discussed below).
At 25°C only
atg-18
RNAi significantly reduced
daf-2(e1368)
mean lifespan, by 15.5% (
p
< 0.0001) (summed data,
Figure 1F
,
Table S2
), a reduction that was significantly greater than in N2 (
p
= 0.0012, CPH, summed data only;
Table S2
). For survival curves comparing RNAi effects of individual
atg
genes on the two genotypes, see
Fig. S2
. In one case,
atg-9
, RNAi slightly increased
daf-2
lifespan (+9.8%,
p
= 0.011, summed data).
The initial tests showing that
bec-1
RNAi reduces
daf-2(e1370)
lifespan were performed at 15°C (
Meléndez et al., 2003
); moreover, a greater N2 life-shortening effect of
bec-1
RNAi at 15°C than 20°C has been reported (
Chen et al., 2019
). Taken together with the weaker RNAi effects at 25°C observed here, this suggested that stronger effects might be seen at 15°C. To explore this we compared effects of
bec-1
RNAi from L4 on N2 and
daf-2(e1370)
at 15°C and 20°C (
N
= 2). However, no suppression of
daf-2(e1370)
Age by
bec-1
RNAi was seen at either temperature (
Figure 1G
;
Table S5
), consistent with findings of an earlier study performed at 15°C (
Hars et al., 2007
) (
Table S1
).
FUDR can alter the effect of
atg
RNAi on lifespan
Since the 1980s FUDR, a thymidylate synthase inhibitor and anti-cancer drug, has sometimes been added to
C. elegans
survival trials to prevent progeny production (
Gandhi et al., 1980
;
Mitchell et al., 1979
). Notably, two previous reports that observed increases in
C. elegans
lifespan given
atg
RNAi employed FUDR. Our own study saw increases in N2 lifespan after
atg-2
and
atg-13
RNAi from L4 with 15 µM FUDR (
Ezcurra et al., 2018
). Another study that saw increases in N2 lifespan given adult-limited
atg-7
,
atg-9
,
bec-1
and
unc-51
RNAi used FUDR at a higher concentration, 800 μM (
Hashimoto et al., 2009
) (E. Nishida, personal communication).
To test for FUDR-dependent effects, we compared the impact of
atg
gene RNAi (
atg-2
,
atg-4.1
,
atg-9
,
atg-13
,
atg-18
and
bec-1
) on N2 lifespan at 20°C with 0, 15 or 800 µM FUDR (
N
= 2). With 0 or 15 µM FUDR, only shortening of lifespan was seen, and FUDR did not significantly alter the effects of RNAi (CPH,
Figure 3A,B
,
Table S6
).
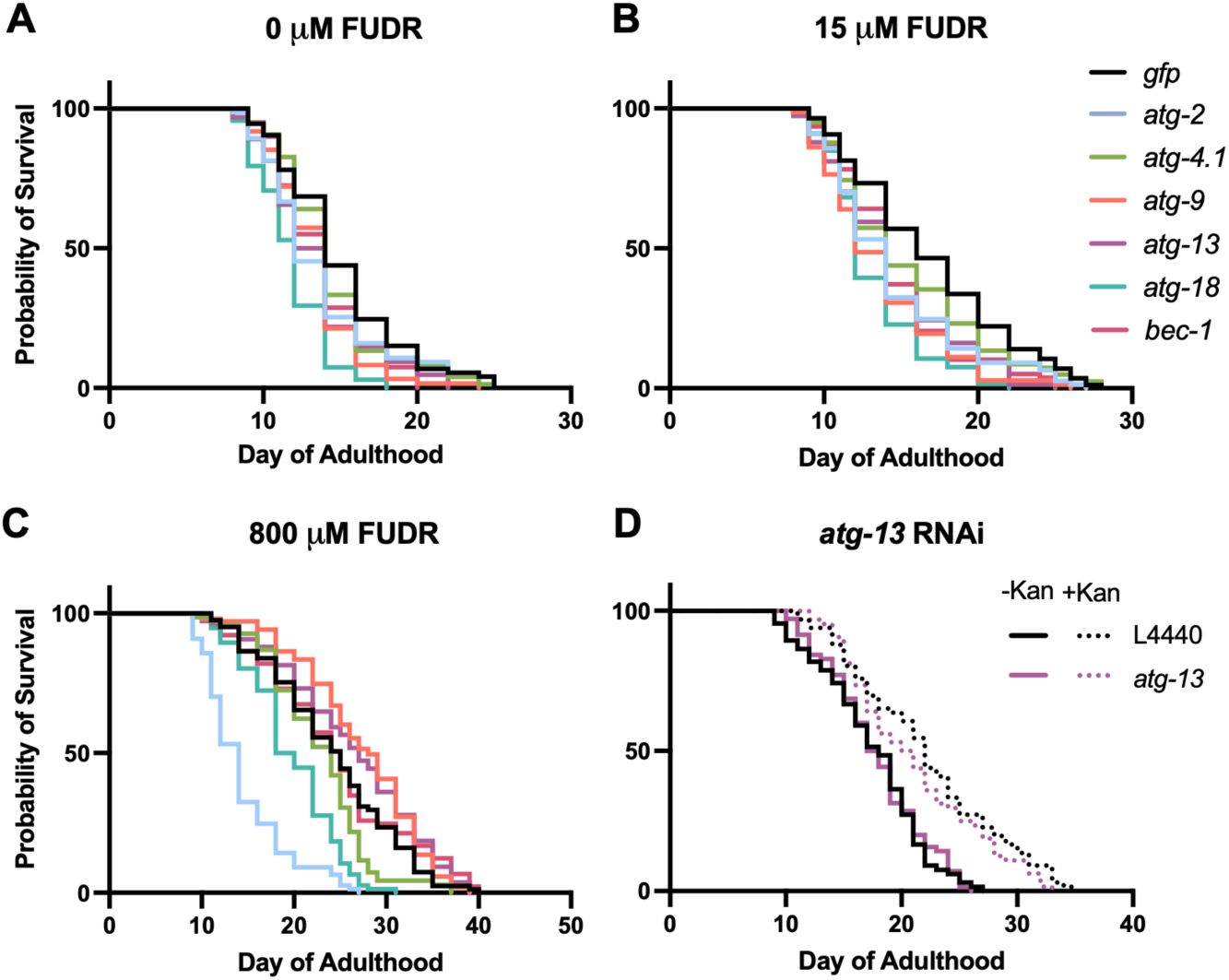
FUDR but not infection alters outcome on
atg
gene RNAi (20°C).
(A-C) Effects of 0 μM, 15 μM and 800 µM FUDR. (D) No alteration by kanamycin of
atg-13
RNAi effect on N2 lifespan. Summed data,
N
= 2; for individual trials and statistical comparisons, see
Table S6
,
S7
.
Addition of 800 µM FUDR increased the lifespan of the
gfp
RNAi negative control by 61.7% (
p
< 0.0001,
Table S6
), perhaps due to prevention of bacterial proliferation, which can otherwise shorten
C. elegans
lifespan (
Garigan et al., 2002
;
Gems and Riddle, 2000
). In the presence of 800 µM FUDR,
atg-9
and
atg-13
RNAi increased lifespan, by +12.5% (
p
= 0.014) and +8.7% (
p
= 0.031), respectively (summed data;
Figure 3C
,
Table S6
). Moreover, the life-shortening effect of
bec-1
RNAi, seen with 0 or 15 µM FUDR, was absent. Again,
atg-18
RNAi robustly reduced lifespan under all three conditions (
Figure 3C
,
Table S6
). The results using 800 µM FUDR are broadly in line with those of
Hashimoto et al. (2009)
, where
atg-9
and
atg-13
RNAi increased lifespan and
atg-18
was one of only 2/14 genes tested where adult-limited RNAi decreased N2 lifespan. This suggests that the increases in lifespan after
atg
RNAi reported in that study could have reflected its use of 800 µM FUDR (
Hashimoto et al., 2009
).
atg
RNAi shortens lifespan in the absence of bacterial infection
Under standard laboratory culture conditions,
C. elegans
lifespan is limited by infection by the
E. coli
food source, such that prevention of bacterial proliferation substantially increases lifespan (
Garigan et al., 2002
;
Gems and Riddle, 2000
). The preceding results could imply that 800 µM FUDR suppresses life-shortening effects of
atg
RNAi by preventing bacterial infection. Xenophagy is generally protective against infection in
C. elegans
(
Balla et al., 2019
;
Jia et al., 2009
). Thus, reduction in lifespan given
atg
gene knockdown could reflect increased susceptibility to infection.
To probe this hypothesis, we compared effects on N2 lifespan of
atg-13
RNAi at 20°C in the absence or presence of the antibiotic kanamycin (Kan), to suppress bacterial infection. In the absence of Kan,
atg-13
RNAi caused a slight reduction in lifespan in these trials that did not reach statistical significance (
Figure 3D
,
Table S7
), in contrast to other trials in this study (
Table S2
,
S6
). Application of Kan extended
C. elegans
lifespan (+27.1%,
p
< 0.0001, summed data,
Table S7
), as previously observed (
Garigan et al., 2002
), and in its presence
atg-13
RNAi resulted in the same slight reduction in lifespan (
Figure 3D
,
Table S7
). These results suggest that life-shortening effects of
atg
RNAi are not solely attributable to increased susceptibility to
E. coli
infection. Moreover, they do not indicate marked condition dependency in
atg
RNAi effects on lifespan with respect to
E. coli
proliferative status.
atg-18
RNAi robustly suppresses
glp-1(e2141)
Age
We next explored more widely the reproducibility and condition dependence of the requirement for autophagy of
C. elegans
longevity. Prevention of germline development by laser surgery or mutation increases lifespan in
C. elegans
hermaphrodites (
Arantes-Oliveira et al., 2002
;
Hsin and Kenyon, 1999
;
Pires da Silva et al., 2024
). Prior tests for possible autophagy dependence of such longevity have largely used the temperature-sensitive
glp-1(e2141)
g
ermline
p
roliferation mutant, which is fertile at 15°C but sterile and with greatly reduced germline development at 25°C. A key study reported strong and reproducible suppression of
glp-1
Age by RNAi of five autophagy-related genes, including
atg-18
and
bec-1
(
Lapierre et al., 2011
) (previous findings summarized in
Table S8
).
We first tested the effect of
atg
RNAi on
glp-1
longevity with animals raised from L1 until L4 stage at 25°C, and maintained at 20°C thereafter, similar to previous studies (
Table S8
). Again, effects of
atg-2
,
atg-4.1, atg-9, atg-13
,
atg-18
and
bec-1
RNA were tested. In the RNAi control
glp-1
increased mean lifespan by +50.9% (
p
< 0.0001, summed data,
Table S9
).
glp-1
lifespan was significantly reduced by
atg-2
,
atg-4.1
,
atg-9
,
atg-18
and
bec-1
RNAi (but not
atg-13
RNAi), and effects were greater in
glp-1
than N2 in all cases except
atg-4.1
(
Figure 4A, B
,
Figure 5
,
Table S9
). However, suppression was only robust (of a large magnitude) for
atg-2
and
atg-18
RNAi (
Figure 4B
,
Figure 5
). Under all six
atg
RNAi conditions,
glp-1
still significantly increased mean lifespan, from +4.12% (
p
= 0.0003) for
atg-18
to +43.8% (
p
< 0.0001) for
atg-4.1
(summed data, 20°C,
Table S9
). This could imply either that
glp-1
Age is not fully autophagy dependent, or that it is but autophagy is not fully suppressed by the RNAi treatment.
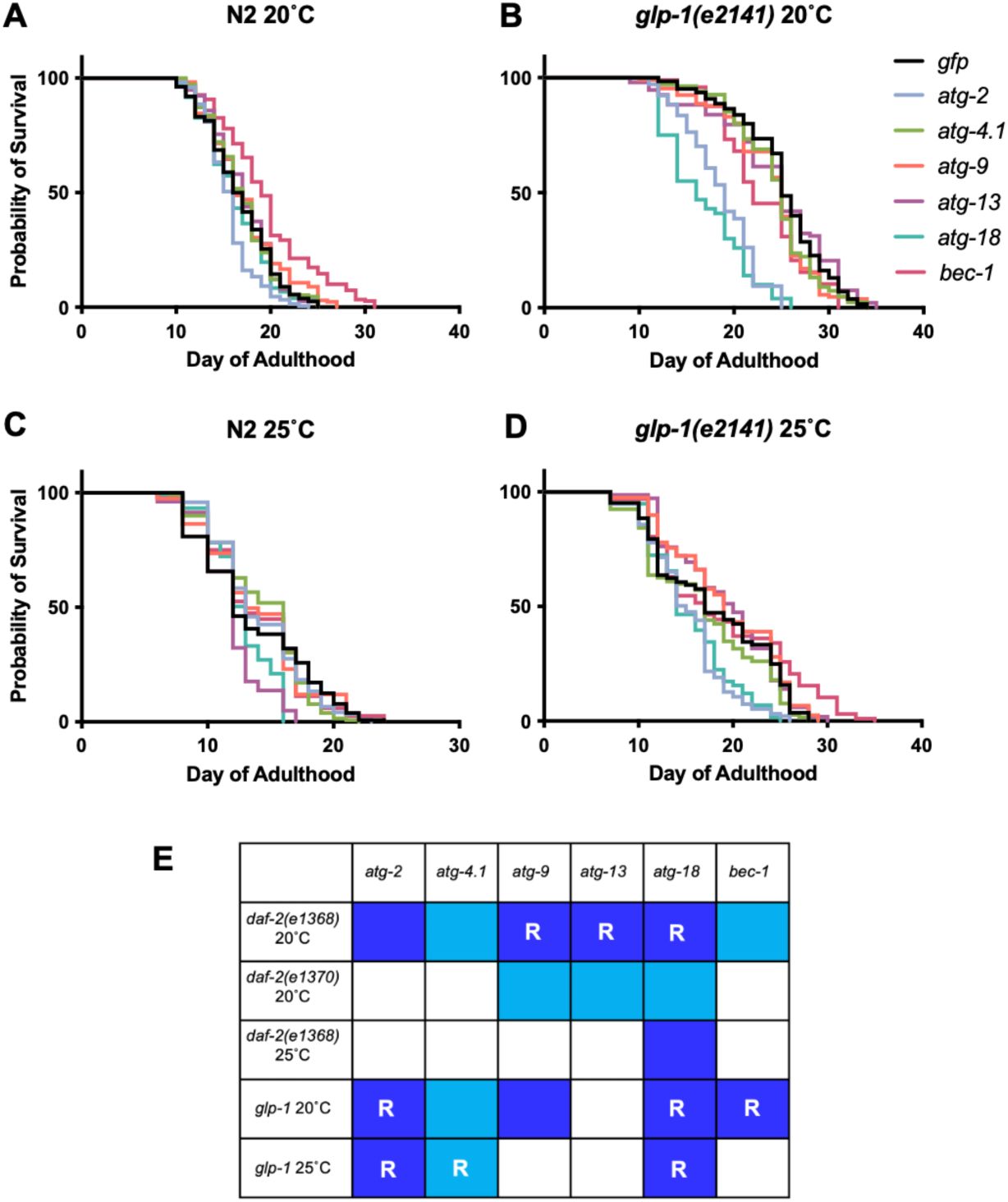
Degree of suppression of
glp-1(e2141)
longevity by
atg
gene RNAi differs greatly between genes.
(A, B) 20°C. (A) Effects on N2. (B) Effects on
glp-1
. (C, D) 25°C. (C) Effects on N2. (D) Effects on
glp-1
. Summed data,
N
= 2-5; for individual trials and statistical comparisons, see
Table S9
. (E) Overview of effects of suppression of Age by RNAi of genes specifying autophagy. Dark blue, reduction of
daf-2
or
glp-1
Age that is significantly greater than in N2. Light blue, reduction of
daf-2
or
glp-1
Age that is not significantly greater than in N2. R, robust suppression, i.e. strong but incomplete suppression of longevity, defined here as <30% increase in mean lifespan in
daf-2
or
glp-1
relative to N2, with RNAi, and as evident from
Figure 2
,
Figure 5
, and
Fig. S2
.
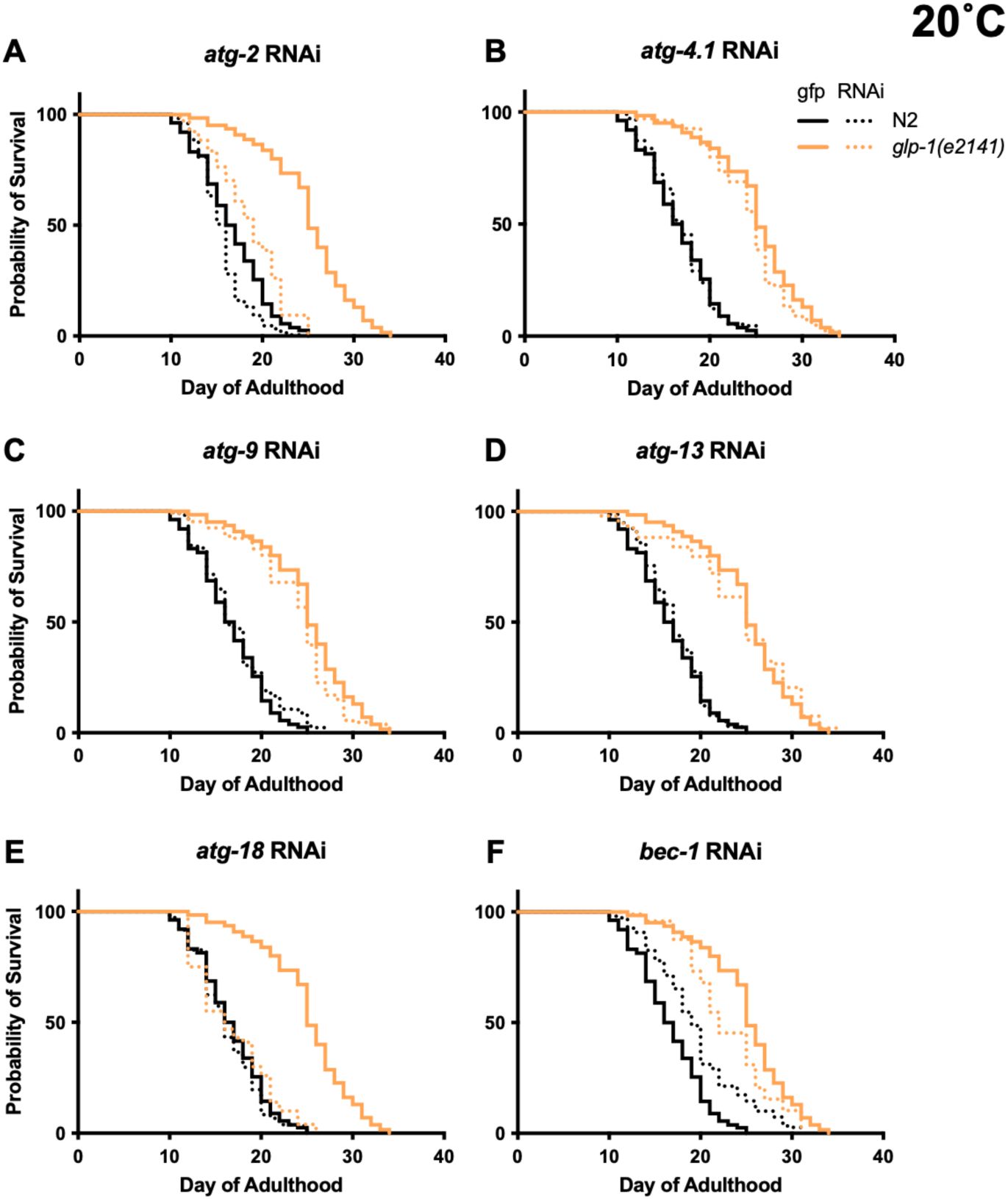
Degree of suppression of
glp-1(e2141)
longevity by
atg
gene RNAi differs greatly between genes (20°C), cf.
Figure 4A,B
(same data).
Summed data,
N
= 2-5; for individual trials and statistical comparisons, see
Table S9
.
In tests with
daf-2(e1368)
life-shortening effects of
atg
RNAi were largely absent at 25°C (apart from
atg-18
) (
Figure 1F
,
Table S2
). We therefore wondered whether temperature might also influence the outcome of
atg
RNAi treatment in
glp-1
mutants. To test this we examined RNAi effects on lifespan at 25°C. Under these conditions,
glp-1
lifespan was significantly reduced by only
atg-2, atg-4.1
, and
atg-18
RNAi (
Figure 4C, D
,
Fig. S3
,
Table S9
), and effects were significantly greater in
glp-1
than N2 only with
atg-2
and
atg-18
RNAi (CPH analysis,
Table S9
). In summary, of the six genes tested only
atg-2
and
atg-18
RNAi robustly suppressed
glp-1
Age, and RNAi effects were weaker at 25°C.
For an overview of the effects of RNAi on
daf-2
and
glp-1
, see
Figure 4E
. In 11 of the 30 conditions tested RNAi knockdown of autophagy caused a greater reduction in lifespan in the long-lived mutant. In 7/30 the RNAi effect was robust, i.e. the mutant longevity was largely suppressed. This suppression was highly condition dependent, differing according to gene knocked down, temperature,
daf-2
allele used, and between
daf-2
and
glp-1
. Notably, only
atg-18
RNAi robustly suppressed longevity in both
daf-2
and
glp-1
mutants.
Why are effects of
atg-18
RNAi stronger than those of other
atg
genes?
The particularly strong life-shortening effects of
atg-18
RNAi could reflect either a greater reduction in autophagy, or the presence in addition to its effects on autophagy of pleiotropic effects not directly related to autophagy. Given that longevity due to either
daf-2
mutation or germline loss are wholly dependent on the FOXO transcription factor DAF-16 (
Hsin and Kenyon, 1999
;
Kenyon et al., 1993
), we wondered whether
atg-18
RNAi might inhibit DAF-16. To probe this two approaches were taken. First, we used a constitutive dauer formation assay. High population density and food depletion causes
C. elegans
larvae to form developmentally arrested dauer larvae (
Cassada and Russell, 1975
).
daf-2
mutants undergo constitutive dauer arrest (the Daf-c phenotype) in a temperature-sensitive manner, and this is fully suppressed by
daf-16
(−) (
Riddle et al., 1981
). However, using a sensitive assay (
daf-2(m41)
, 22°C, giving a mix of dauer and non dauers), we detected no reduction in the number of dauers formed given
atg
gene RNAi (
atg-2
,
atg-13
and
atg-18
) (
Fig. S4A
).
Second, we tested a GFP reporter for a gene whose expression is elevated in
daf-2
mutants in a
daf-16
-dependent manner (
ftn-1
), using strains previously constructed (
Ackerman and Gems, 2012
). GFP levels were compared in
daf-2(m577)
and
daf-16(mgDf50)
;
daf-2
backgrounds. As expected, GFP levels were higher in
daf-2
than in
daf-16; daf-2
(
Fig. S4B
). RNAi did not suppress the
daf-2
-induced increase of
ftn-1::gfp
expression (
Fig. S4B
). These findings argue against a pleiotropic effect of
atg-18
on DAF-16 function.
Inhibiting autophagy does not reduce vitellogenin accumulation
Finally, we further investigated the hypothesis that autophagy promotes biomass repurposing in
C. elegans
. Inhibition of yolk synthesis or of autophagy delays intestinal atrophy and yolk pool accumulation, suggesting that intestinal biomass is repurposed for yolk synthesis (
Benedetto and Gems, 2019
;
Ezcurra et al., 2018
;
Sornda et al., 2019
). In principle, this could involve repurposing into yolk protein (vitellogenin) or yolk lipid. To test the former possibility, wild-type hermaphrodites or
fog-2(q71)
(feminization
o
f
g
ermline) mutant females were subjected to
atg
RNAi. The
fog-2
mutant, which lacks self-sperm and so lays no eggs, was used to avoid possible effects of
atg
RNAi on fertility, reduction of which can increase vitellogenin levels within nematodes (
Sornda et al., 2019
).
For none of the six
atg
genes tested did RNAi detectably reduce yolk protein levels, either in N2 hermaphrodites or
fog-2
females (
Figure 6
). This implies that autophagic machinery, including that in the intestine, does not enhance yolk protein production. Thus, if intestinal biomass repurposing occurs, then it likely supports yolk secretion or yolk lipid production.
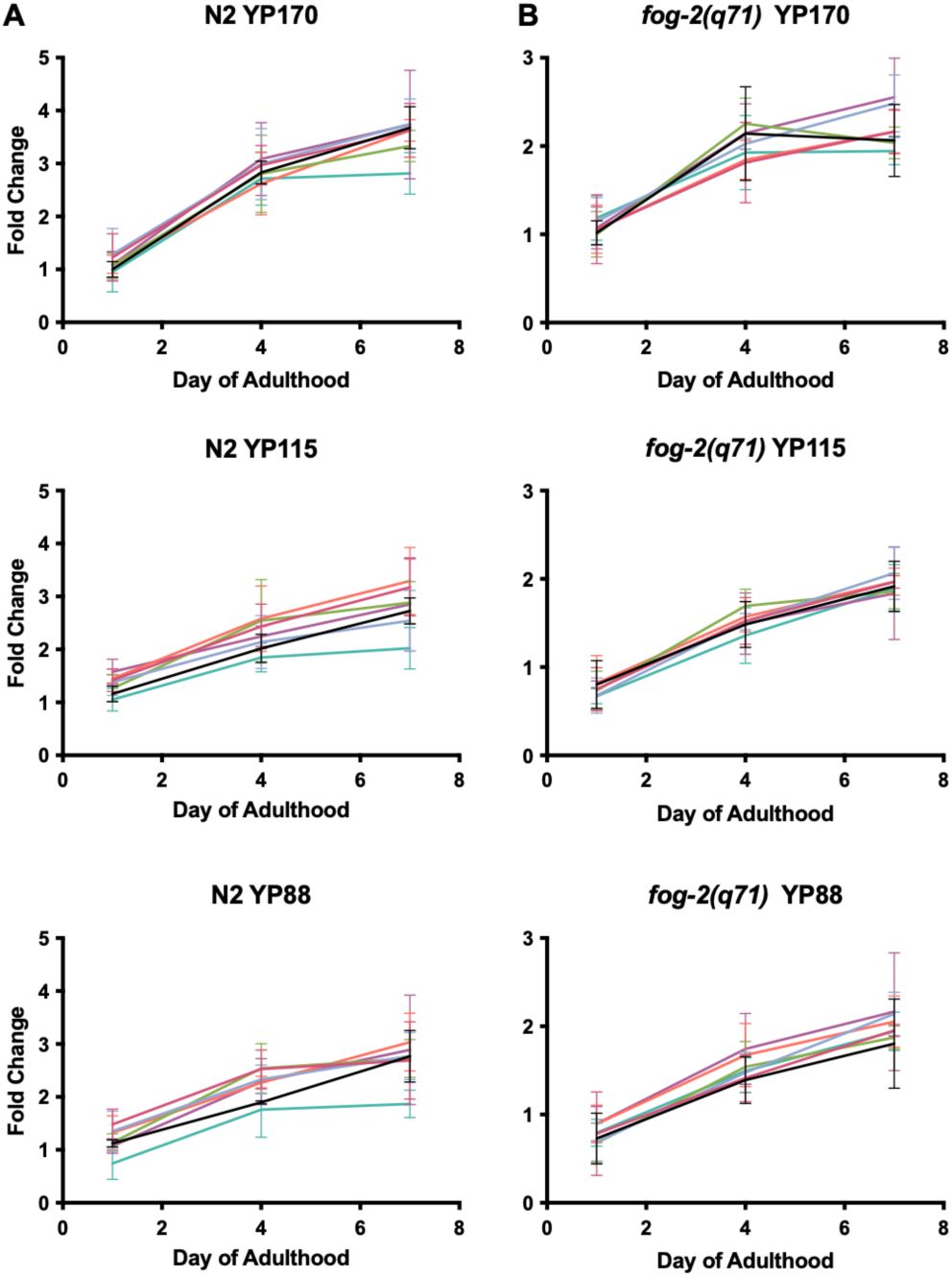
Absence of effect of
atg
RNAi on vitellogenin accumulation.
Fold change of yolk proteins YP170, YP115 and YP88 in N2 and
fog-2
normalized to
gfp
day 1 (
N
= 3). In no case is vitellogenin level significantly different to that in the
gfp
RNAi control at any time point (two-way ANOVA,
Table S10
). For raw data for vitellogenin levels see
Supplementary Dataset 3
.
Discussion
Overall, the effects of reducing
atg
gene function described here are ambiguous with respect to the role of autophagy in
daf-2
or
glp-1
longevity, neither strongly supporting or excluding it. However, they are consistent with a role of autophagy in the longevity of weaker, class 1
daf-2
alleles at lower temperatures (
Figure 1C,D,F,G
,
Table S2
). Class 1 allele-limited suppression of
daf-2
Age has been seen previously, for example in epistasis tests with
daf-12
(encoding a dafachronic acid receptor) (
Gems et al., 1998
;
Larsen et al., 1995
) and
skn-1
(Nrf2-like transcription factor) (
Tullet et al., 2008
). This could reflect a role of autophagy in longevity assurance limited to conditions of mild IIS reduction, or sensitivity to differential effects of
daf-2
on distinct downstream signaling outputs, such as phosphatidylinositol 3-kinase and Ras signaling (
Patel et al., 2008
).
atg
RNAi suppresses
daf-2(e1368)
Age at 20°C but not 25°C (
Figure 1C,F
,
Table S2
). Given that for hypomorphic
daf-2
alleles (such as
e1368
and
e1370
) many mutant traits, including Age, show some degree of temperature sensitivity (
Gems et al., 1998
;
Riddle and Albert, 1997
), the absence of suppression at 25°C may reflect the increased severity of the
daf-2(e1368)
mutant phenotype at this higher temperature. However, life-shortening effects of
atg
RNAi on N2 are also reduced at 25°C, suggesting that additional mechanisms may also mediate such temperature sensitivity.
Resolving discrepancies between past studies of autophagy dependence of
daf-2
Age
The increasing non-reproducibility of many experimental findings, that has been referred to as the
reproducibility crisis
, is a problem that particularly afflicts biological and biomedical research (
Baker, 2016
;
Lithgow et al., 2017
;
Prinz et al., 2011
;
Ritchie, 2020
;
Voelkl et al., 2020
). A strength of
C. elegans
as an experimental model is the relative ease with which such discrepancies can be resolved, at least in principle. This is thanks to the use of standardized culture conditions across the
C. elegans
research community, and of nematode strains based on the same isogenic wild-type strain (N2), plus the relatively low cost and short duration of experiments.
Regarding lifespan assays in particular, possible reasons for discrepant findings include clearly identifiable differences in experimental design, such as use or not of FUDR. Less obvious causes include cryptic variation in genetic background (
Zhao et al., 2019
), or subtle differences in culture conditions (e.g. due to batch variation in Bacto Peptone, a constituent of nematode growth medium) (
Petrascheck, 2014
).
Our findings potentially resolve several discrepancies between earlier studies relating to the possible role of autophagy in
daf-2
mutant longevity. Previous studies found that inhibiting autophagy either suppressed or enhanced
daf-2
Age, or had no effect (
Table S1
). One study showing suppression used the weak class 1 allele
daf-2(mu150)
(
Hansen et al., 2008
). This is consistent with our observation that,
atg-18
aside,
atg
RNAi can suppress Age in a class 1 but not a class 2 allele (
Figure 1C,D
Table S2
).
Does
bec-1
RNAi suppress
daf-2(e1370)
Age? The initial test suggesting this was performed at 15°C, and employed RNAi by injection of the mothers of assayed individuals (
Meléndez et al., 2003
). In a subsequent study at 15°C, in which both mothers and adult progeny were exposed to
bec-1
RNAi by feeding,
daf-2(e1370)
lifespan appeared not to be shortened (
Hars et al., 2007
). Several further trials under different conditions did not observe a life-shortening effect either (
Hashimoto et al., 2009
;
Toth et al., 2008
). In the present study, we saw no shortening of
daf-2(e1370)
lifespan in summed data with adult-limited
bec-1
RNAi by feeding, at either 15°C or 20°C (
Figure 1D,G
,
Fig. S3
,
Table S2
,
S5
). Taken together with earlier evidence, our observations suggest that suppression of
daf-2(e1370)
Age by
bec-1
RNAi is not a readily reproducible finding.
Several studies reported lifespan extension following
atg
RNAi (
Ezcurra et al., 2018
;
Hashimoto et al., 2009
). Here we were able to reproduce this effect for several
atg
genes by applying high dose FUDR (800 µM), thus recapitulating findings by
Hashimoto et al (2009)
, and potentially accounting for the life span increases seen in that study; it was only subsequent to that study that evidence emerged of the capacity of FUDR to alter effects of interventions that impact lifespan (
Aitlhadj and Sturzenbaum, 2010
;
Anderson et al., 2016
;
Burnaevskiy et al., 2018
;
Van Raamsdonk and Hekimi, 2011
;
Zhao et al., 2019
).
Regarding issues with replicating published findings, in an earlier study we observed increases in N2 lifespan given
atg-2
and
atg-13
RNAi (15 µM FUDR present) (
Ezcurra et al., 2018
). However, this finding proved not to be robust to replication (
Figure 3B
,
Table S6
), perhaps reflecting inherent effect variability, condition dependence or a combination of the two. This again illustrates the value, as with the effects of
bec-1
RNAi on
daf-2(e1370)
Age, of repeated verification of effects of interventions on
C. elegans
lifespan.
As with
daf-2
, our tests with the
glp-1
germline proliferation mutant did not clearly support the view that mutant longevity is autophagy dependent. However, consistent with an earlier study (
Lapierre et al., 2011
), we observed that
atg-18
and
bec-1
RNAi reduce
glp-1
lifespan more than in wild-type at 20°C.
Why does
atg-18
RNAi more strongly suppress
daf-2
Age?
Of the six autophagy-determining genes tested here, RNAi of
atg-18
showed a greater capacity to suppress both
daf-2
and
glp-1
Age than the other five.
atg-18
appears to have replaced
bec-1
as the gene of choice for autophagy-related epistasis studies in
C. elegans
(
Chang et al., 2017
;
Minnerly et al., 2017
). Notably,
atg-18
was one of only 2/14 autophagy genes tested where RNAi during development caused a high level (>50%) of larval growth arrest or lethality (
Hashimoto et al., 2009
). The more marked effects of
atg-18
RNAi could reflect either greater inhibition of autophagy, or pleiotropy in which processes other than autophagy are altered.
Regarding pleiotropy, several proteins in the canonical autophagy pathway have recently been found to participate in other processes. For example, ATG8 (LGG-1 in
C. elegans
) functions in trafficking of single-membrane organelles (
Nieto-Torres et al., 2021
), and
C. elegans
ATG-16.2 contributes to neuronal exopher formation via its WD40 domain (
Yang et al., 2024
). In an as yet unexplained instance of
atg
gene idiosyncrasy, mutation of
atg-16.2
,
atg-18
or
bec-1
retards the cell cycle in
C. elegans
germline cells, while that of
atg-7
does not (
Ames et al., 2017
).
Concerning possible
atg-18
pleiotropy, several studies suggest this possibility. ATG-18 is a predicted WIPI (WD repeat protein interacting with phosphoinositides) family member. In humans there are four WIPI proteins, WIPI1 - WIPI4;
C. elegans atg-18
is more closely related to WIPI1/WIPI2, while
epg-6
more closely resembles WIPI3/WIPI4 (
Lu et al., 2011
). Notably, deletion mutations of
atg-18
and
epg-6
appear to cause similar reductions in levels of autophagy, yet only
atg-18
shortens lifespan (
Takacs et al., 2019
).
Next, a
C. elegans
studies of rescue of longevity and fat storage in
daf-2(e1370); atg-18(gk378)
double mutants by tissue-specific expression of
atg-18
(+) revealed a major role of
atg-18
in food-sensing chemosensory neurons, potentially reflecting a vesicle trafficking role in neurosecretion (possibly of neuropeptides) (
Jia et al., 2019
;
Minnerly et al., 2017
). Whether such a role is unique to
atg-18
, or shared by other
atg
genes remains unclear.
Third, recent evidence suggests that ATG-18 activates HLH-30/TFEB (transcription factor EB). HLH-30 is a master transcriptional regulator of autophagy and lysosomal biogenesis, that becomes nuclear localized in
daf-2(e1370)
and
glp-1(e2141)
mutants, and whose inhibition reduces longevity in both contexts (
Lapierre et al., 2013
;
Lin et al., 2018
;
Wong et al., 2023
). Notably, among genes up-regulated upon over-expression of
atg-18
, ones with HLH-30 elements in their promoters are over-represented, and similarly in genes down-regulated in an
atg-18(gk378)
mutant. Also, heat shock-induced nuclear localization of HLH-30 is suppressed by
atg-18
RNAi, as is longevity resulting from
atg-18
over-expression (
Schmauck-Medina et al., 2026
).
Inhibition of HLH-30 by
atg-18
RNAi could potentially explain its unusually strong life-shortening effects. If correct, this could reflect greater suppression of autophagy, but also, in principle, other effects of HLH-30 inhibition. HLH-30 affects a variety of traits, including adult reproductive diapause (Gerisch et al., 2020), proteostasis (
Shalash et al., 2025
), sex differences in immunity (
Sohn et al., 2025
), and resistance to various insults, including infectious pathogens (
El-Houjeiri et al., 2019
;
Visvikis et al., 2014
;
Wani et al., 2021
), starvation (
Harvald et al., 2017
;
O’Rourke and Ruvkun, 2013
;
Settembre et al., 2013
), and oxidative and heat stress (
Lin et al., 2018
).
While in at least some of these cases autophagy likely contributes to
hlh-30
-mediated effects, given that this transcription factor also promotes expression of many genes unrelated to autophagy (
Chen et al., 2017
;
Lin et al., 2018
;
Shalash et al., 2025
;
Sohn et al., 2025
;
Visvikis et al., 2014
), other functions may also be involved. For example, in response to infection HLH-30 can activate expression of signaling (including IIS), autophagy-related and immunity-related genes, and both of latter can contribute to infection resistance (
Chen et al., 2017
;
Sohn et al., 2025
;
Visvikis et al., 2014
). In conclusion, if the more severe effects of
atg-18
RNAi on lifespan reported here are attributable to reduced HLH-30 activity, this could well reflect greater reduction of autophagy. However, it remains possible that other functions controlled by HLH-30 also play a role, e.g. relating to immunity (
Chen et al., 2017
;
Visvikis et al., 2014
) and protein quality control (
Shalash et al., 2025
). Also notable is that HLH-30 regulates gene expression combinatorially with DAF-16 (
Lin et al., 2018
), mutation of which fully suppresses
daf-2(rf)
longevity (
Kenyon et al., 1993
).
Autophagy in biomass repurposing during programmatic aging
Autophagic processes play a major role in tissue degeneration related to biomass repurposing in semelparous organisms (that reproduce once and then die), particularly plants (
Avila-Ospina et al., 2014
;
Gems et al., 2021
). Previous studies support the hypothesis that intestinal biomass is repurposed for synthesis of yolk that, subsequent to egg laying, is vented to support larval development (
Kern et al., 2021
), and that autophagy facilitates this biomass conversion (
Ezcurra et al., 2018
;
Sornda et al., 2019
). If late-life mortality is promoted by intestinal atrophy, then preventing it should extend lifespan. Consistent with this, intestinal atrophy and yolk production are suppressed in
daf-2
mutants (
Depina et al., 2011
;
Ezcurra et al., 2018
), and blocking yolk production both retards intestinal atrophy and modestly increases lifespan (
Ezcurra et al., 2018
;
Murphy et al., 2003
;
Sornda et al., 2019
). This could imply that inhibiting autophagy, by retarding intestinal atrophy could extend lifespan under some conditions, and several earlier studies observed life extension after
atg
RNAi (
Ezcurra et al., 2018
;
Hashimoto et al., 2009
;
Wilhelm et al., 2017
).
The results of the present study are in line with the view that intestinal atrophy, though a salient feature of
C. elegans
senescence, by itself contributes only weakly to late-life mortality. Consistent with this, inhibition of
daf-2
using auxin-induced degradation of DAF-2 protein can strongly increasing lifespan even when initiated only at advanced ages, long after intestinal atrophy has occurred, and without any detectable reversal of major senescent pathologies (
Molière et al., 2024
;
Venz et al., 2021
). One possibility is that
atg
RNAi has antagonistic effects on late-life mortality: modestly reducing it due to suppression of intestinal atrophy (as seen when vitellogenin synthesis is inhibited) (
Ezcurra et al., 2018
;
Murphy et al., 2003
;
Sornda et al., 2019
), but also increasing it, due to disruption of essential cellular functions.
The gut-to-yolk biomass repurposing hypothesis drew particularly on the observation that blocking yolk protein (vitellogenin) synthesis or autophagy reduced intestinal atrophy and pseudocoelomic yolk pool size (
Ezcurra et al., 2018
;
Sornda et al., 2019
); given that yolk pools contain yolk protein and lipid (
Ezcurra et al., 2018
;
Garigan et al., 2002
;
Herndon et al., 2002
;
Palikaras et al., 2017
), such putative biomass repurposing could increase production of yolk protein, yolk lipid or both. Here we tested whether
atg
RNAi reduces yolk protein levels, and found that it did not (
Figure 6
). If the hypothesis is correct, then this would support the view that promotion by autophagy of repurposing of intestinal biomass into yolk involves yolk lipid rather than yolk protein, or overall yolk secretion. In principle, this could involve mass export of stored lipid, either by means of lipophagy, or perhaps secretory autophagy (
Ponpuak et al., 2015
). As an aside, we note that there is evidence that vitellogenin levels can affect autophagy (
Seah et al., 2016
).
Concluding remarks
This study reassesses the claim that
daf-2
Age is autophagy dependent. While the results do not entirely prove or disprove this claim, they clearly show that evidence of dependence varies greatly with context, and suggest that autophagy only contributes substantially to
daf-2
Age given a weak reduction in IIS.
One limitation in establishing the role of autophagy in
daf-2
mutant longevity is the difficulty of reliably measuring its level of activity. While several assays using fluorescent reporters to identify putative autophagosomes suggest possible increases in autophagy levels (
Chang et al., 2017
;
Hansen et al., 2008
;
Meléndez et al., 2003
),
daf-2
mutants also show marked reductions in both protein synthesis rate (
Depuydt et al., 2013
) and turnover rate (
Dhondt et al., 2016
;
Visscher et al., 2016
), including reduced ATG-18 turnover rate (
Visscher et al., 2016
). This is more consistent with a reduction rather than an increase in autophagy, and in line with observed suppression of intestinal atrophy in
daf-2
mutants (
Ezcurra et al., 2018
).
One issue raised by condition dependence of a given effect is that it creates a risk of condition selection bias, where choice of experimental conditions can bias the outcome of an experiment. Where condition dependence has been identified, condition selection bias may be avoided by selection of multiple conditions to test a given hypothesis. In the present case, this could entail comparing effects on
daf-2(e1368)
and
daf-2(e1370)
at 20°C, ideally without FUDR use, and RNAi tests with multiple
atg
genes, at least until the reason for the idiosyncratic effects of
atg-18
RNAi are understood. General questions relating to how to deal with condition dependency have been discussed previously (
Munafò and Davey Smith, 2018
;
Voelkl et al., 2020
).
A second, wider issue is that condition dependence risks generating a situation where different research articles make conflicting claims, supporting different views held by different groups of researchers. For science to progress effectively it is necessary for research communities to resolve discrepancies between published findings, though the work required can be tedious, a task akin to washing the dishes in a communal household. It requires identifying and flagging findings that are either condition dependent or, seemingly, unreproducible and, where possible, distinguishing the two. For
C. elegans
at least, such a tidying process is highly feasible, as is use of methodologies devised to reduce such discrepancies (
Driscoll et al., 2025
;
Lucanic et al., 2017
;
Petrascheck and Miller, 2017
), which is a particular virtue of this model organism.
Materials and Methods
Culture methods and strains
C. elegans
maintenance was performed using standard protocols (
Brenner, 1974
). Unless otherwise stated, all strains were on nematode growth media (NGM, containing Bacto Peptone) with plates seeded with
E. coli
OP50 to provide a food source. An N2 hermaphrodite stock recently obtained from the Caenorhabditis Genetics Center was used as wild type (N2H) (
Zhao et al., 2019
). Genotypes of most mutants used are as described in Wormbase (
www.wormbase.org
). Strains used included CB4027
glp-1(e2141)
, GA633
daf-2(m577)
;
wuIs177 [Pftn-1::gfp lin-15(+)]
, GA643
daf-16(mgDf50)
;
daf-2(m577)
;
wuIs177 [Pftn-1::gfp lin-15(+)]
], GA1930
daf-2(e1370)
, GA1945
daf-2(m41)
, GA1960
daf-2(e1368)
, and JK574
fog-2(q71) V
.
Constitutive dauer larva formation assay
daf-2(m41)
animals were maintained on RNAi feeding strains for at least two generations prior to analysis. For the dauer larva formation assay, performed at 22°C, 10 L4-stage hermaphrodites were transferred onto RNAi plates and allowed to lay eggs for approximately 6 h, after which adults were removed. Dauers were scored at 166 h.
Epifluorescence microscopy
Nematodes were anaesthetized with 10 μl 2 mM levamisole on 2% agar pads prior to imaging. For imaging, we used either a Zeiss Axioskop 2 plus microscope with Hamamatsu ORCA-ER digital camera C4742-95 and Volocity 6.3 software (Macintosh version) for image acquisition; or an ApoTome.2 Zeiss microscope with a Hamamatsu digital camera C13440 ORCA-Flash4.0 V3 and Zen software for image acquisition.
RNA-mediated interference (RNAi)
RNAi by feeding was performed using RNAi plasmids transformed into
E. coli
OP50(xu363) as previously described (
Xiao et al., 2015
). Bacterial transformants were selected on LB agar plates containing 10 µg/ml tetracycline and 25 µg/ml carbenicillin. Inserts of all RNAi feeding clones were confirmed by sequencing. Origins of plasmids in RNAi feeding strains:
atg-13
,
bec-1
: Ahringer library (
Kamath et al., 2003
);
atg-2
,
atg-4.1
,
atg-9
: Vidal library (
Rual et al., 2004
). As an
atg-18
RNAi feeding clone was not available in local RNAi libraries, a custom
atg-18
RNAi plasmid was generated by cloning the target sequence into the L4440 vector using primers
gcctccacttcctgttgaag
and
gagactcttttcgtcggca
.
For RNAi induction, bacteria were grown for 16 h at 37°C with shaking (200 rpm) in 5 ml LB supplemented with 25 µg/ml carbenicillin. Overnight cultures were diluted 1:100 into fresh LB containing 25 µg/ml carbenicillin and grown for a further 4 h or until reaching an OD₆₀₀ of ∼0.4. Expression of double-stranded RNA was induced by addition of 1 mM IPTG, and cultures were incubated for an additional 1 h. Following induction, cultures were allowed to cool to room temperature, concentrated five-fold by centrifugation, and seeded onto NGM plates supplemented with 25 µg/ml carbenicillin and 1 mM IPTG. Seeded plates were allowed to dry at room temperature before use.
To minimize IPTG degradation, RNAi plates were stored in foil-lined boxes. Typically, unseeded plates were stored at 4°C for up to one month, while seeded plates were used within two weeks. RNAi treatment was initiated from the L4 larval stage unless otherwise stated.
Survival analysis
Nematodes were maintained at a density of 25-30 per plate, and transferred daily during the egg laying period, and every 6-7 days thereafter. The L4 stage was defined as day 0. Mortality was scored every 1-2 days, with worms scored as alive if they showed any movement, either spontaneously or in response to gentle touch with a worm pick.
RNA extraction, cDNA synthesis and RT-qPCR
Approximately 100–200 day 4 adult animals per treatment group were lysed in 250 µl of TRIzol™ reagent by vortexing for 10 min at 4°C, followed by a 10 min incubation on ice; this cycle was repeated three times. RNA was purified using the RNeasy Mini Kit (QIAGEN, Cat. No. 74104) according to the manufacturer’s instructions, including on-column DNase digestion using the RNase-Free DNase Set (QIAGEN, Cat. No. 79256). cDNA was synthesized from 100 ng of total RNA using the SuperScript™ First-Strand Synthesis System (Thermo Fisher Scientific, Cat. No. 10684803).
PCR primers were designed using Primer-BLAST with the following parameters: maximum primer length of 20 nucleotides, PCR product size of 70–200 bp, and exon–exon spanning where possible. Primer efficiency and R² values were determined from standard curves to ensure suitability for quantitative analysis and comparable amplification efficiencies between primer pairs. Primers were used at a final concentration of 2 µM.
Quantitative PCR was performed using Fast SYBR™ Green chemistry on a QuantStudio™ 6 Flex Real-Time PCR System (Applied Biosystems) with 384-well plates and a total reaction volume of 10 µl. Cycling conditions were 95°C for 20 s, followed by 40 cycles of 95°C for 1 s and 65°C for 20 s, with fluorescence data acquisition during the annealing/extension step. Melt curve analysis was performed following amplification to confirm product specificity. A full list of primers used in this study is provided in
Table S11
. mRNA levels were normalized to mRNA levels from two housekeeping genes,
cdc-42
and
pmp-3
(
Hoogewijs et al., 2008
).
Electrophoresis of
C. elegans
yolk protein
N2 and
fog-2(q71)
worms were synchronized by performing an egg lay and allowing nematodes to grow at 20°C until they reached the L4 stage. L4 worms were then transferred to fresh seeded RNAi plates and maintained under standard culture conditions at 20°C. Five worms were harvested at day 1, day 4 and day 7 into microcentrifuge Eppendorf tubes containing 10 μl M9 buffer. Before running the gel, 10 μl of 2x Laemmli sample buffer was added. Samples were incubated at 70°C and vortexed periodically for 15 min. Samples were then incubated at 95°C and vortexed periodically for 5 min, and were centrifuged at 6,000 rpm for 15 min.
10 μl of sample was loaded into wells of Invitrogen NuPAGE Bis-Tris protein gels. 5 μl of CozyHi pre-stained protein ladder was loaded at the left side of the gels. The running buffer used was 5% XT MOPS (Bio-Rad). The gels were then run at 150V for 2 hr. Gels were removed from cassette and placed in 100 ml of Coomassie staining solution overnight. Gels were then washed with distilled water three times, then placed into destaining solution and soaked for 40 min. Gels were then washed with distilled water and stored at 4°C until imaged. Gels were imaged and saved as 8 bit grayscale TIF files. Images of gels were analysed using Fiji software. Bands of interest (e.g. myosin, YP170, YP115, YP88) were selected manually based on their molecular weight, and intensity of the bands was measured and exported to Microsoft Excel for further analysis.
Data analysis and statistics
Data were plotted using ePrism 9.0 (GraphPad Software, Boston, MA, USA) or Jupyter Notebook using Python with the matplotlib, pandas and NumPy libraries. Statistical tests were performed on raw data using either Prism 9 or JMP Pro 15 (JMP Statistical Discovery LLC, Cary, NC, USA) unless otherwise stated. The specific tests and post hoc corrections performed are described in the figure legends. To detect alterations in lifespan, the log rank test was used. To compare the magnitude of changes in lifespan, Cox Proportional Hazard (CPH) analysis was used. RT-qPCR data was analyzed using the ΔΔCt method and t-tests. To compare yolk protein levels, a two-way ANOVA was used. For lifespan trials, no statistical methods were used to predetermine sample size. The experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment.
Data availability
The datasets used and/or analyzed during the current study are available with this article, and also from the corresponding author Prof. David Gems (david.gems@ucl.ac.uk) on request.
Acknowledgements
We thank Georg Fuellen (Rostock University), Kailiang Jia (Florida Atlantic University), Alicia Meléndez (Queens College-CUNY), Eisuke Nishida (RIKEN), and John Labbadia, Om Patange and Hongyuan Wang (UCL) for useful discussion, and/or comments on the manuscript, and Minh Tran Dang, Anna Girtle, Changtai Li, Gadea Meecham-Garcia and Suzie Mishima for minor research contributions. Some strains were provided by the Caenorhabditis Genetics Center, which is funded by NIH Office of Research Infrastructure Programs (P40 OD010440).
Additional information
Funding
This work was supported by a Wellcome Trust Investigator Award (215574/Z/19/Z) to D.G..
Funding
Wellcome Trust (WT)
https://doi.org/10.35802/215574
David Gems
Additional files
Supplementary tables 2-7, 9-11
References
Insulin/IGF-1 and hypoxia signaling act in concert to regulate iron homeostasis in C. elegans
PLoS Genet
8
:e1002498
Google Scholar
The use of FUdR can cause prolonged longevity in mutant nematodes
Mech Ageing Dev
131
:364–5
Google Scholar
Autophagy in healthy aging and disease
Nat Aging
1
:634–650
Google Scholar
A non-cell-autonomous role of BEC-1/BECN1/Beclin1 in coordinating cell-cycle progression and stem cell proliferation during germline development
Curr Biol
27
:905–913
Google Scholar
C. elegans lifespan extension by osmotic stress requires FUdR, base excision repair, FOXO, and sirtuins
Mech Ageing Dev
154
:30–42
Google Scholar
Regulation of life-span by germ-line stem cells in Caenorhabditis elegans
Science
295
:502–505
Google Scholar
Conceptual Breakthroughs in The Evolutionary Biology of Aging
Academic Press
Google Scholar
Autophagy, plant senescence, and nutrient recycling
J Exp Botany
65
:3799–3811
Google Scholar
1,500 scientists lift the lid on reproducibility
Nature
533
:452–4
Google Scholar
Natural variation in the roles of C. elegans autophagy components during microsporidia infection
PLoS One
14
:e0216011
Google Scholar
The free radical theory of aging matures
Physiol Rev
78
:547–581
Google Scholar
Autophagy promotes visceral aging in wild-type C. elegans
Autophagy
15
:731–732
Google Scholar
Aging and immortality: quasi-programmed senescence and its pharmacologic inhibition
Cell Cycle
5
:2087–102
Google Scholar
The genetics of Caenorhabditis elegans
Genetics
77
:71–94
Google Scholar
Reactivation of RNA metabolism underlies somatic restoration after adult reproductive diapause in C. elegans
eLife
7
:e36194
https://doi.org/10.7554/eLife.36194
Google Scholar
The dauerlarva, a post-embryonic developmental variant of the nematode Caenorhabditis elegans
Dev Biol
46
:326–342
Google Scholar
Spatiotemporal regulation of autophagy during Caenorhabditis elegans aging
eLife
6
:e18459
https://doi.org/10.7554/eLife.18459
Google Scholar
HLH-30/TFEB-mediated autophagy functions in a cell-autonomous manner for epithelium intrinsic cellular defense against bacterial pore-forming toxin in C. elegans
Autophagy
13
:371–385
Google Scholar
Adiponectin receptor PAQR-2 signaling senses low temperature to promote C. elegans longevity by regulating autophagy
Nat Commun
10
:2602
Google Scholar
Genomes optimize reproduction: aging as a consequence of the developmental program
Physiology
20
:252–259
Google Scholar
Regulation of Caenorhabditis elegans vitellogenesis by DAF-2/IIS through separable transcriptional and posttranscriptional mechanisms
BMC Physiol
11
:11
Google Scholar
Reduced insulin/IGF-1 signaling and dietary restriction inhibit translation but preserve muscle mass in Caenorhabditis elegans
Mol Cell Proteomics
12
:3624–3639
Google Scholar
FOXO/DAF-16 activation slows down turnover of the majority of proteins in C. elegans
Cell Rep
16
:3028–3040
Google Scholar
NIA Caenorhabditis Intervention Testing Program: identification of robust and reproducible pharmacological interventions that promote longevity across experimentally accessible, genetically diverse populations
Geroscience
Google Scholar
The Transcription factors TFEB and TFE3 link the FLCN-AMPK signaling axis to innate immune response and pathogen resistance
Cell Rep
26
:3613–3628
Google Scholar
C. elegans eats its own intestine to make yolk leading to multiple senescent pathologies
Curr Biol
28
:2544–2556
Google Scholar
A simple method for maintaining large, aging populations of Caenorhabditis elegans
Mech Ageing Dev
12
:137–150
Google Scholar
Genetic analysis of tissue aging in Caenorhabditis elegans: a role for heat-shock factor and bacterial proliferation
Genetics
161
:1101–1112
Google Scholar
Understanding hyperfunction: an emerging paradigm for the biology of aging
Ageing Res Rev
74
:101557
Google Scholar
Alternative perspectives on aging in C. elegans: reactive oxygen species or hyperfunction?
Antioxid Redox Signal
19
:321–329
Google Scholar
Antioxidant defense and aging in C. elegans: is the oxidative damage theory of aging wrong?
Cell Cycle
8
:1681–7
Google Scholar
Biological constraint, evolutionary spandrels and antagonistic pleiotropy
Ageing Res Rev
101
:102527
Google Scholar
Reproductive suicide: similar mechanisms of aging in C. elegans and Pacific salmon
Front Cell Dev Biol
9
:688788
Google Scholar
Genetic, behavioral and environmental determinants of male longevity in Caenorhabditis elegans
Genetics
154
:1597–1610
Google Scholar
Two pleiotropic classes of daf-2 mutation affect larval arrest, adult behavior, reproduction and longevity in Caenorhabditis elegans
Genetics
150
:129–155
Google Scholar
A role for autophagy in the extension of lifespan by dietary restriction in C. elegans
PLoS Genet
4
:e24
Google Scholar
Autophagy as a promoter of longevity: insights from model organisms
Nat Rev Mol Cell Biol
19
:579–593
Google Scholar
Aging: A theory based on free radical and radiation chemistry
J Gerontol
11
:298–300
Google Scholar
Autophagy regulates ageing in C. elegans
Autophagy
3
:93–5
Google Scholar
Multi-omics analyses of starvation responses reveal a central role for lipoprotein metabolism in acute starvation survival in C. elegans
Cell Syst
5
:38–52
Google Scholar
Lifespan extension by suppression of autophagy genes in Caenorhabditis elegans
Genes Cells
14
:717–726
Google Scholar
Stochastic and genetic factors influence tissue-specific decline in ageing C. elegans
Nature
419
:808–814
Google Scholar
Selection and validation of a set of reliable reference genes for quantitative sod gene expression analysis in C. elegans
BMC Mol Biol
9
:9
Google Scholar
Signals from the reproductive system regulate the lifespan of C. elegans
Nature
399
:362–366
Google Scholar
Autophagy genes protect against Salmonella typhimurium infection and mediate insulin signaling-regulated pathogen resistance
Proc Natl Acad Sci U S A
106
:14564–9
Google Scholar
Neuroendocrine regulation of fat metabolism by autophagy gene atg-18 in C. elegans dauer larvae
FEBS Open Bio
9
:1623–1631
Google Scholar
Systematic functional analysis of the Caenorhabditis elegans genome using RNAi
Nature
421
:231–237
Google Scholar
A C. elegans mutant that lives twice as long as wild type
Nature
366
:461–464
Google Scholar
C. elegans ageing is accelerated by a self-destructive reproductive programme
Nat Commun
14
:4381
Google Scholar
C. elegans feed yolk to their young in a form of primitive lactation
Nat Commun
12
:5801
Google Scholar
The TFEB orthologue HLH-30 regulates autophagy and modulates longevity in Caenorhabditis elegans
Nat Commun
4
:2267
Google Scholar
Autophagy and lipid metabolism coordinately modulate life span in germline-less C. elegans
Curr Biol
21
:1507–1514
Google Scholar
Genes that regulate both development and longevity in Caenorhabditis elegans
Genetics
139
:1567–1583
Google Scholar
DAF-16/FOXO and HLH-30/TFEB function as combinatorial transcription factors to promote stress resistance and longevity
Nat Commun.
9
:4400
Google Scholar
A long journey to reproducible results
Nature
548
:387–388
Google Scholar
The WD40 repeat PtdIns(3)P-binding protein EPG-6 regulates progression of omegasomes to autophagosomes
Dev Cell
21
:343–57
Google Scholar
Impact of genetic background and experimental reproducibility on identifying chemical compounds with robust longevity effects
Nat Commun
8
:14256
Google Scholar
Evolution of ageing as a tangle of trade-offs: energy versus function
Proc Biol Sci
286
:20191604
Google Scholar
Autophagy genes are essential for dauer development and life-span extension in C. elegans
Science
301
:1387–1391
Google Scholar
The cell non-autonomous function of ATG-18 is essential for neuroendocrine regulation of Caenorhabditis elegans lifespan
PLoS Genet
13
:e1006764
Google Scholar
Synchronous growth and aging of Caenorhabditis elegans in the presence of fluorodeoxyuridine
J Gerontol
34
:28–36
Google Scholar
Improved resilience and proteostasis mediate longevity upon DAF-2 degradation in old age
Geroscience
46
:5015–5036
Google Scholar
Robust research needs many lines of evidence
Nature
553
:399–401
Google Scholar
How We Age: The Science of Longevity
Princeton and Oxford
:
Princeton University Press
Google Scholar
Genes that act downstream of DAF-16 to influence the lifespan of C. elegans
Nature
424
:277–284
Google Scholar
Beyond autophagy: the expanding roles of ATG8 proteins
Trends Biochem Sci
46
:673–686
Google Scholar
MXL-3 and HLH-30 transcriptionally link lipolysis and autophagy to nutrient availability
Nat Cell Biol
15
:668–76
Google Scholar
Ectopic fat deposition contributes to age-associated pathology in Caenorhabditis elegans
J Lipid Res
58
:72–80
Google Scholar
Clustering of genetically defined allele classes in the Caenorhabditis elegans DAF-2 insulin/IGF-1 receptor
Genetics
178
:931–46
Google Scholar
Is the oxidative stress theory of aging dead?
Biochim Biophys Acta
1790
:1005–14
Google Scholar
Carnivorous elegans
Worm Breeder’s Gazette
20
Google Scholar
Computational analysis of lifespan experiment reproducibility
Front Genet
8
:92
Google Scholar
Decoding lifespan secrets: the role of the gonad in Caenorhabditis elegans aging
Front Aging
5
:1380016
Google Scholar
Secretory autophagy
Curr Opin Cell Biol
35
:106–116
Google Scholar
Believe it or not: how much can we rely on published data on potential drug targets?
Nat Rev Drug Discov
10
:712
Google Scholar
Genetic and environmental regulation of dauer larva development
In:
Riddle DL
Blumenthal T
Meyer BJ
et al.
, editors.
C. elegans II
Plainview, N.Y.
:
Cold Spring Harbor Laboratory Press
pp. 739–768
Google Scholar
Interacting genes in nematode dauer larva formation
Nature
290
:668–671
Google Scholar
Science Fictions: Exposing Fraud, Bias, Negligence and Hype in Science
London, UK
:
Vintage (Penguin Random House)
Google Scholar
Toward improving Caenorhabditis elegans phenome mapping with an ORFeome-based RNAi library
Genome Res
14
:2162–8
Google Scholar
ATG-18/WIPI2 drives longevity in an HLH-30/TFEB-dependent manner
bioRxiv
Google Scholar
Autophagy-mediated longevity is modulated by lipoprotein biogenesis
Autophagy
12
:261–272
Google Scholar
TFEB controls cellular lipid metabolism through a starvation-induced autoregulatory loop
Nat Cell Biol
15
:647–58
Google Scholar
HLH-30/TFEB rewires the chaperone network to promote proteostasis upon perturbations to the coenzyme A and iron-sulfur cluster biosynthesis pathways
Aging Cell
24
:e70038
Google Scholar
Beneficial and detrimental effects of reactive oxygen species on lifespan: a comprehensive review of comparative and experimental studies
Front Cell Dev Biol
9
:628157
Google Scholar
A cytoprotective perspective on longevity regulation
Trends Cell Biol
23
:409–420
Google Scholar
HLH-30/TFEB mediates sexual dimorphism in immunity in Caenorhabditis elegans
Autophagy
21
:283–297
Google Scholar
Production of YP170 vitellogenins promotes intestinal senescence in C. elegans
J Gerontol A
74
:1180–1188
Google Scholar
On the nature of the aging process
Proc Natl Acad Sci U S A
45
:30–45
Google Scholar
ATG-18 and EPG-6 are both required for autophagy but differentially contribute to lifespan control in Caenorhabditis elegans
Cells
8
:236
Google Scholar
Longevity pathways converge on autophagy genes to regulate life span in Caenorhabditis elegans
Autophagy
4
:330–8
Google Scholar
Direct inhibition of the longevity-promoting factor SKN-1 by insulin-like signaling in C. elegans
Cell
132
:1025–38
Google Scholar
FUdR causes a twofold increase in the lifespan of the mitochondrial mutant gas-1
Mech Ageing Dev
132
:519–521
Google Scholar
End-of-life targeted degradation of DAF-2 insulin/IGF-1 receptor promotes longevity free from growth-related pathologies
eLife
10
:e71335
https://doi.org/10.7554/eLife.71335
Google Scholar
Proteome-wide changes in protein turnover rates in C. elegans models of longevity and age-related disease
Cell Rep
16
:3041–3051
Google Scholar
Innate host defense requires TFEB-mediated transcription of cytoprotective and antimicrobial genes
Immunity
40
:896–909
Google Scholar
Reproducibility of animal research in light of biological variation
Nat Rev Neurosci
21
:384–393
Google Scholar
Regulation of Caenorhabditis elegans RNA interference by the daf-2 insulin stress and longevity signaling pathway
Cold Spring Harb Symp Quant Biol
69
:429–31
Google Scholar
NHR-49/PPAR-α and HLH-30/TFEB cooperate for C. elegans host defense via a flavin-containing monooxygenase
eLife
10
Google Scholar
Neuronal inhibition of the autophagy nucleation complex extends life span in post-reproductive C. elegans
Genes Develop
31
:1561–1572
Google Scholar
Pleiotropy, natural selection and the evolution of senescence
Evolution
11
:398–411
Google Scholar
Neuronal HLH-30/TFEB modulates peripheral mitochondrial fragmentation to improve thermoresistance in Caenorhabditis elegans
Aging Cell
22
:e13741
Google Scholar
RNAi interrogation of dietary modulation of development, metabolism, behavior, and aging in C. elegans
Cell Rep
11
:1123–1133
Google Scholar
Autophagy protein ATG-16.2 and its WD40 domain mediate the beneficial effects of inhibiting early-acting autophagy genes in C. elegans neurons
Nat Aging
4
:198–212
Google Scholar
A fln-2 mutation affects lethal pathology and lifespan in C. elegans
Nat Commun
10
:5087
Google Scholar
Article and author information
Author information
Cite all versions
You can cite all versions using the DOI
https://doi.org/
10.7554/eLife.110620
. This DOI represents all versions, and will always resolve to the latest one.
Copyright
© 2026,
Hsiung et al.
This article is distributed under the terms of the
Creative Commons Attribution License
, which permits unrestricted use and redistribution provided that the original author and source are credited.
Metrics
views
0
downloads
0
citations
0
Views, downloads and citations are aggregated across all versions of this paper published by eLife.

